* This is the latest installment of “Problematica.” It is written by Max Dresow (gloriously unemployed, but soon to be affiliated with the University of Copenhagen) and Katherine Valde (Loyola University)…
Anomalocaris reared up, mouth open, its spiny arms outstretched to their furthest limit. It was preparing to fall on a diminutive “lace crab” (Marrella splendens), which was paddling like hell to get away. In front of Marrella were plastered the words “Evolution’s Big Bang” in giant, boldface letters. It was all taking place on the cover of TIME magazine (December 4, 1995), whose cover story promised to illuminate how “life as we know it appeared in an amazing biological frenzy that changed the planet almost overnight.”
The year 1995 was a remarkable time in Cambrian studies. Long a theater of intense debate about pretty much everything, a fragile consensus had begun to form about the timing and “explosiveness” of the Cambrian radiation. Life on earth originated around four billion years ago, and for the next three-and-a-half billion years remained mostly confined to the microbial grade. “Then, 543 million years ago…, within the span of no more than 10 million years, creatures with teeth and tentacles and claws and jaws materialized with the suddenness of apparitions.” This was the “Cambrian explosion,” as described by TIME magazine’s J. Madeleine Nash, and in 1995, the once-controversial idea was becoming mainstream.
Within a year, however, the “Cambrian explosion hypothesis” was again embroiled in controversy. What happened in the interim was the publication of the first “molecular clock” study to use DNA sequence data to constrain the age of major groups.* This found that “invertebrates [i.e., protostomes] diverged from chordates [deuterostomes] about a billion years ago, [or] twice as long ago as the Cambrian” (Wray et al. 1996, 568). The date was admittedly not very precise; yet it seemed to refute the contemporary estimate— put forward by James Valentine and colleagues— that the ancestor of protostomes and deuterostomes lived perhaps thirty million years before the Cambrian, or about 575 million years ago (Valentine et al. 1996).
[* A “molecular clock” is a method for inferring the age of clades (or, if you prefer, lineage-splitting events), which works on the assumption that molecular sequences change at predictable rates over long timescales.]
Thus (re-) commenced a heated debate in paleontology between advocates of a fairly literal reading of the fossil record and advocates of a long cryptic history for major animal groups. The debate is still unresolved (and so, for that matter, is a parallel debate about the origin of angiosperms, or flowering plants). And yet recently— with virtually no hoopla— researchers took a major step towards its resolution. Part of this was technical, and had to do with refinements in molecular clock dating techniques, together with better data. But the more interesting part was philosophical, and concerned a new approach to uncertainty, as well as, perhaps, a new “philosophy of molecular clock dating.”
We will attempt to characterize this “philosophy” in a moment. First, however, it will be useful to fill in some of the history, which explains why the traditional attitude of paleontologists towards molecular clocks has been one of distrust, and even antagonism.
Molecular clocks and the Cambrian explosion: a history of conflict
The first molecular clock study to investigate the origin of animals was conducted by a paleontologist, Bruce Runnegar. Published in 1982, it used differences in the amino acid sequences of alpha- and beta-hemoglobin proteins to date “the initial radiation of animal phyla”— as it happens, to about a billion years ago (Runnegar 1982, 14). The study was criticized on methodological grounds, and failed to have much impact on the course of debates about animal origins (e.g., Erwin 1989). However, in 1996, a molecular clock study appeared that “set the paleontological world on its ear” (Knoll 2003, 200).
The study emerged from a collaboration between Gregory Wray and Leo Shapiro— both evolutionary biologists— and Jeffrey Levinton— a paleontologist. It concluded, in line with Runnegar’s estimate, that the major animal phyla had emerged between 1.0 and 1.2 billion years ago, or “about twice as long ago as the Cambrian.” Unlike Runnegar’s study it used DNA sequence data to build the clock, sourced from several independent gene families. Thus, when it concluded that animals had passed nearly half their history without leaving a single unambiguous trace in the rocks, people noticed.
Early molecular clock studies worked on the assumption that molecular sequences change at predictable rates over long timescales. So, given a certain amount of differentiation between the “same” gene or molecule in different lineages, an investigator could use predetermined rates of sequence change to estimate when the lineages diverged from a common ancestor. Provided that rates of sequence divergence had been properly measured, and provided these rates could be extrapolated across the tree, the method was expected to return at least a reliable ballpark estimate of the true age of a group.
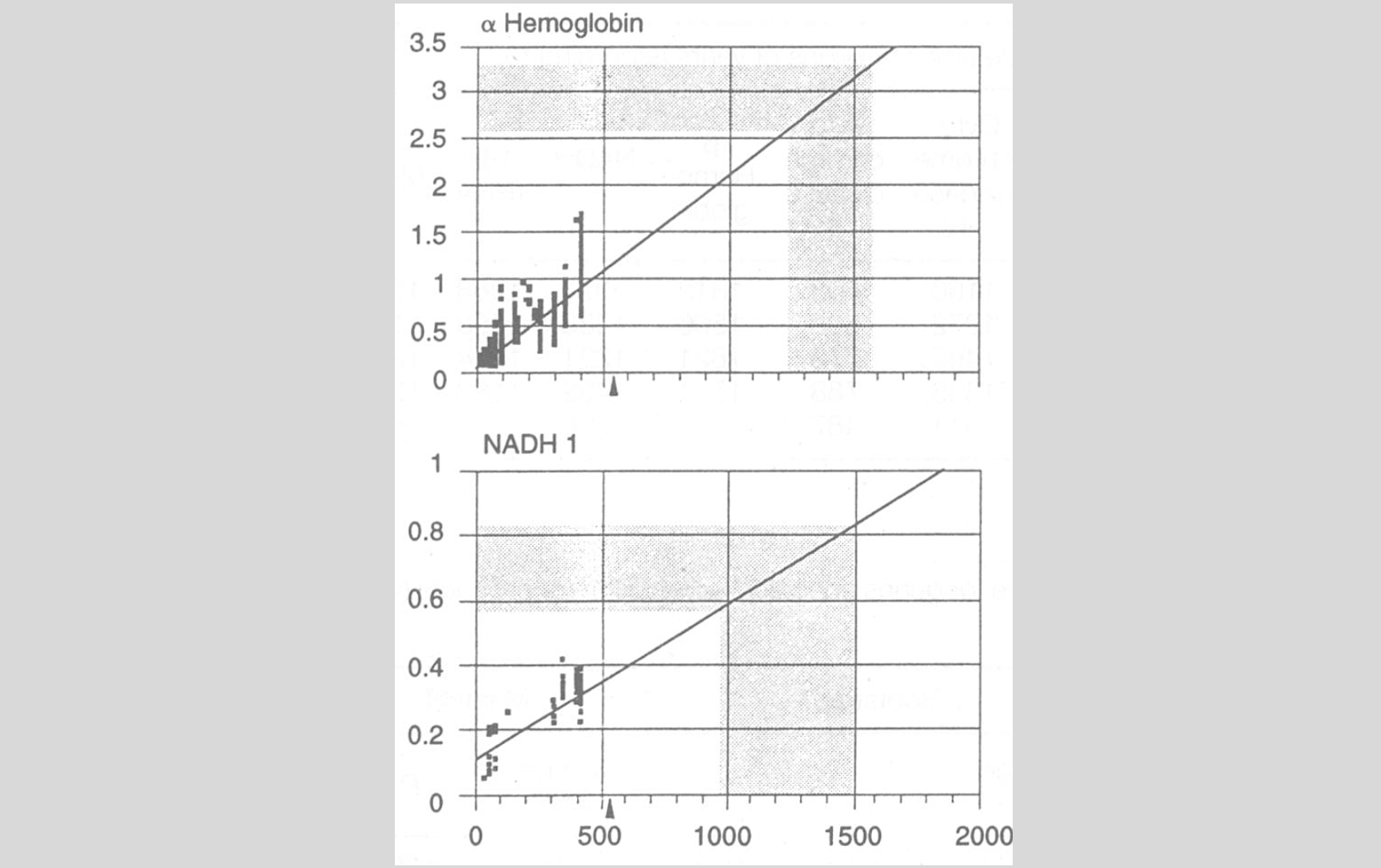
To build the 1996 clock, Wray and colleagues began by gathering nucleotide sequence data for “homologous” genes in different lineages. Then, they calculated the amount of divergence between pairs of lineages and plotted this against the estimated time of divergence for the two lineages based on the fossil record (see the figure, below). They used vertebrate genes to calibrate the clock, since the vertebrate fossil record was assumed to be pretty good. Anyway, it enabled them to characterize rates of molecular change for each gene that could then be extended to parts of the tree where the fossil record was missing or equivocal.
Sequence divergence rates and estimated interphylum divergence times, from Wray et al. (1996). The reproduced graphic shows two genes, alpha-hemoglobin and NADH 1. Diverge times (in Ma) are plotted on the x-axis; genetic distance is plotted on the y-axis. The data points come from vertebrate taxa with high-quality fossil records. The shaded parts of the plots indicate the full range of “invertebrate-vertebrate genetic distances” and the implied range of divergence times
Wray and colleagues were interested in one divergence event in particular: the split between protostomes and deuterostomes. To that end, they measured sequence differences between a large number of protostome and deuterostome species, and— using the rates measured on vertebrate materials— produced a large number of splitting dates (one per species-pair, per gene). What they found was that “all mean divergence time estimates… substantially predate[d] the Cambrian,” with mean divergence times between protostomes and deuterostomes averaging between 1.0 and 1.2 billion years ago (Wray et al. 1996, 570). The average concealed much variability between the “clocks.” Genes coding for hemoglobin suggested the split happened about 1.6 billion years ago, for instance, whereas genes coding for cytochrome oxidase suggested a date of just 800 million years. Still, all of these were “incompatible with the Cambrian explosion hypothesis of rapid, shallow interphylum distances.” Not a year after the TIME magazine cover, notions of explosive evolution in the Cambrian were again on the hot seat.
Many paleontologists were immediately suspicious of the new dates. Asked for a comment, Andy Knoll of Harvard quipped that “molecular clocks are not Timexes.” Others shared his caution, and contributed a measure of indignation on behalf of the embattled fossil record (see Benton 2009). The fossil record was incomplete, sure— but could it possibly be this incomplete? For many paleontologists, the idea that animals could have existed for hundreds of millions of years without making a peep was quite literally incredible.
The technical features of Wray and colleagues’ study soon came in for criticism. The study used too few genes to produce reliable estimates, critics said. More seriously, the genes they did use exhibited variability in substitution rate (meaning that some clocks ticked faster than others). Newer analyses, including one published in 1998, revised age-estimates toward conformity with the fossil record (Ayala et al. 1998). But the 1998 study placed the protostome-deuterostome split at about 670 million years ago (Ma), leaving a hundred-million-year gap between the time of the split and the first ambiguous glimmerings of the animal fossil record. Other studies favored older dates, with estimates reduced by perhaps 20–30% (e.g., Bromham et al. 1998, Wang et al. 1999). So the debate remained polarized.
Around 2008, a group led by Doug Erwin and Kevin Peterson set out to produce the definitive molecular clock study of early animal evolution. To do this, they used “relaxed” clock methods (designed to accommodate fluctuations in substitution rate) to estimate divergence times for more than a hundred species spread across the tree. What they found was that the last common ancestor of all living animals probably lived around 800 Ma, with the first “crown groups” appearing a hundred million years later (Erwin et al. 2011).* The major pulse of bilaterian divergence took place in the Ediacaran Period (then thought to stretch from 635 to about 541 Ma). As for bilaterian crown groups, these appeared between the end of the Ediacaran and the end of the Cambrian, in agreement with the fossil record of well-skeletonized taxa.
[* The “crown group” of a clade consists of the last common ancestor of the living members of the clade together with all its descendants, living and extinct. The “stem group,” by contrast, consists of all the lineages that branched beneath the base of the crown group and above the last common ancestor of the crown group and its closest living relative.]
The conclusion was apparently unavoidable. There had been a lag, perhaps as long as 200 million years, between the origin of animals and their debut in the fossil record. Any account of the Cambrian explosion would have to come to grips with it, or else show that the clock had been improperly wound.
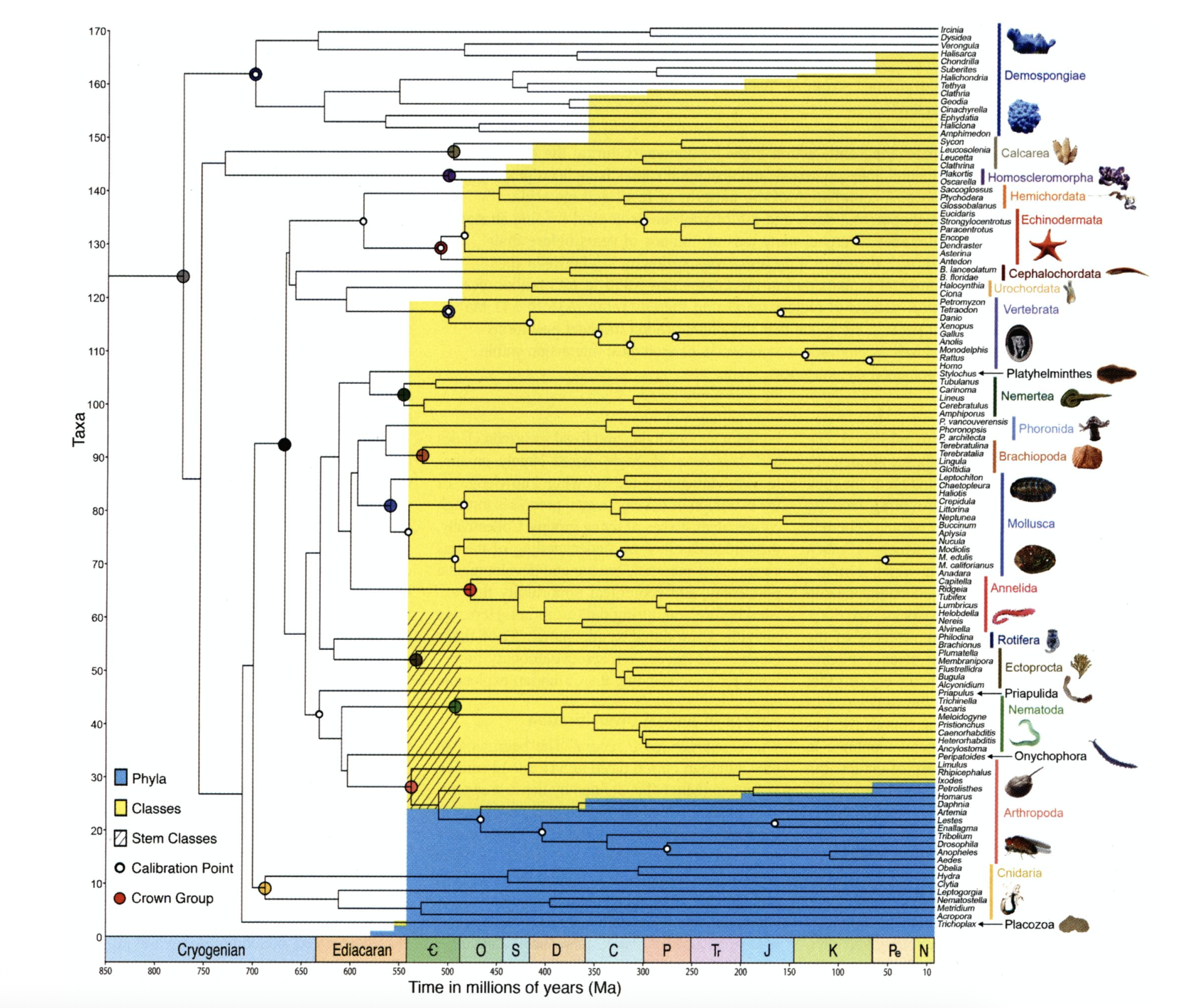
The results of Erwin et al.’s (2011) molecular clock study, showing the estimated divergence times of major clades, superimposed on a colored graphic showing the total number of phyla and classes known from the fossil record of each interval
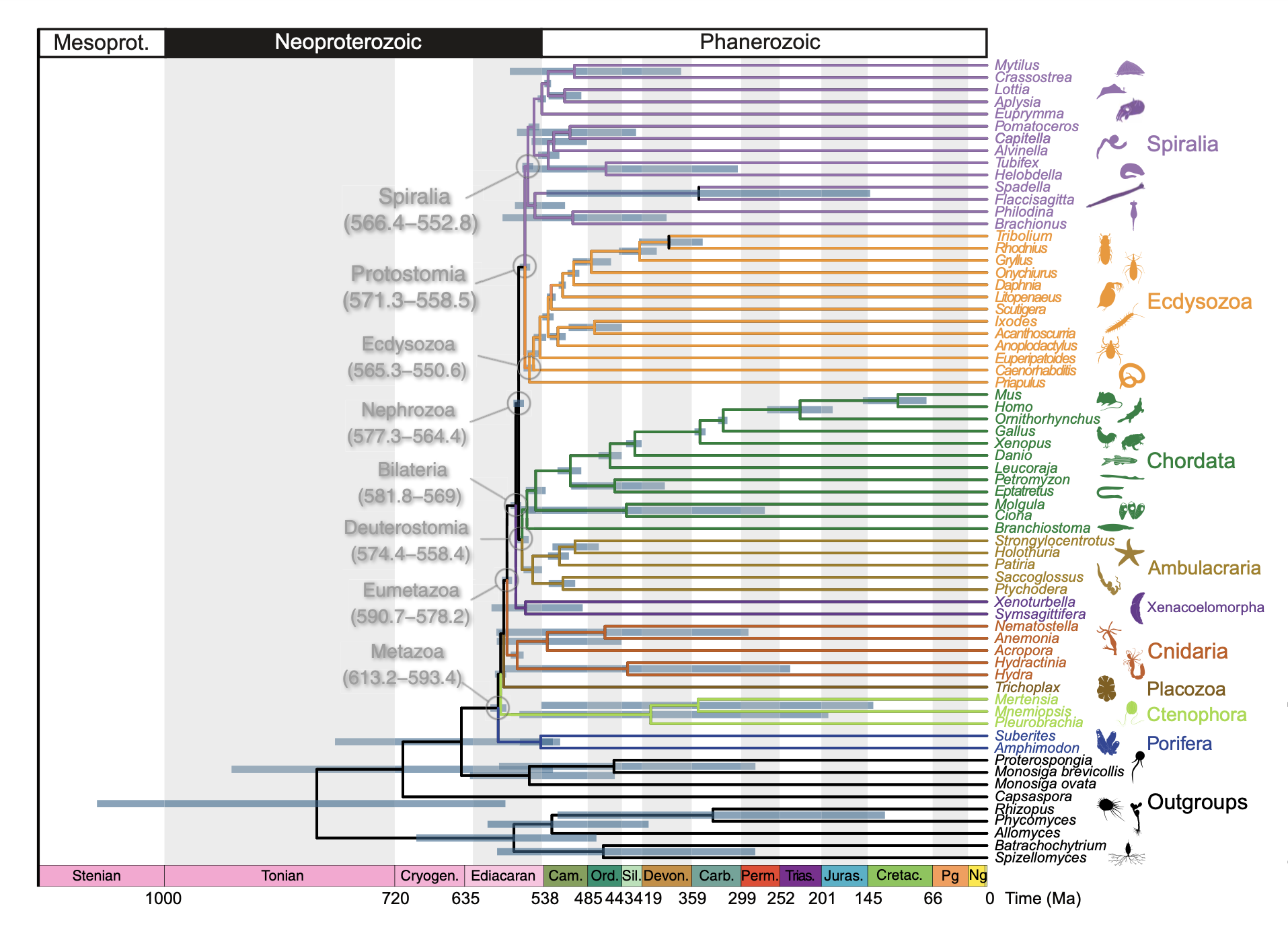
The result stood for over a decade. Subsequent analyses, using more extensive datasets, only strengthened it, even pushing the origin of bilaterians to a time before the last snowball glaciation (>720 Mya) (Dohrmann and Wörheide 2017). It came to seem unlikely that the result was going anywhere: paleontologists were going to have to accept a sizeable gap between the origin of major clades and their first appearance in the fossil record. Yet the most recent molecular clock study of early animal evolution, published in 2024, came to a radically different conclusion (Carlisle et al. 2024). According to this study, animals originated in the early Ediacaran, between 613 and 593 million years ago. Bilaterians originated later, between about 580 and 569 million years ago, and protostomes and deuterostomes later still, between about 575 and 558 million years ago. These are significantly earlier dates than Erwin and company produced— around 200 million years earlier for the origin of animals— implying missing fossil records of just 20 to 40 million years for major groups.
So, what gives?
All about uncertainty
To begin, there were some methodological differences between the studies. The Erwin et al. study employed concatenation, for example, treating multiple loci as a single nonrecombining locus (and so, ignoring gene tree discordance) (Tiley et al. 2020). It also employed a more limited dataset, which did not sample the ctenophores, and only sampled three of the four classes of Porifera, or sponges (Dohrmann and Wörheide 2017). However, by far the most important difference concerned the handling of fossil evidence, and what came to the same thing, the attitude the investigators took toward uncertainty.
Backing up a moment: popular “Bayesian” molecular clock methods (employed by Erwin et al. and Carlisle et al.) require the user to specify both minimum and maximum constraints on the ages of branch points, or “nodes.”* These correspond to the lower and upper limits of admissible ages for those nodes, respectively. Minimum constraints are generally based on the oldest known crown group representative of the clade in question (since the clade cannot be younger than its oldest fossilized member). Maximum constraints, by contrast, “attempt to constrain how much older the clade may be… [based] on qualified negative evidence,” like the evidence of sister taxa in sediments that would be expected to preserve members of the clade in question had they been present (Carlisle et al. 2024, 2). As Carlisle and colleagues observe, “maxima are often arbitrary but objective, in the sense that they are defined on the basis of a well-studied and readily dated deposit.” Still, they have often been defined conservatively, indicating that a large envelope of uncertainty surrounds the age of key nodes, which then propagates through the rest of the analysis to produce inflated divergence estimates (relative to the estimates produced by less conservative maxima).
[* For a primer on Bayesian methods of molecular clock dating, see dos Reis et al. (2015).]
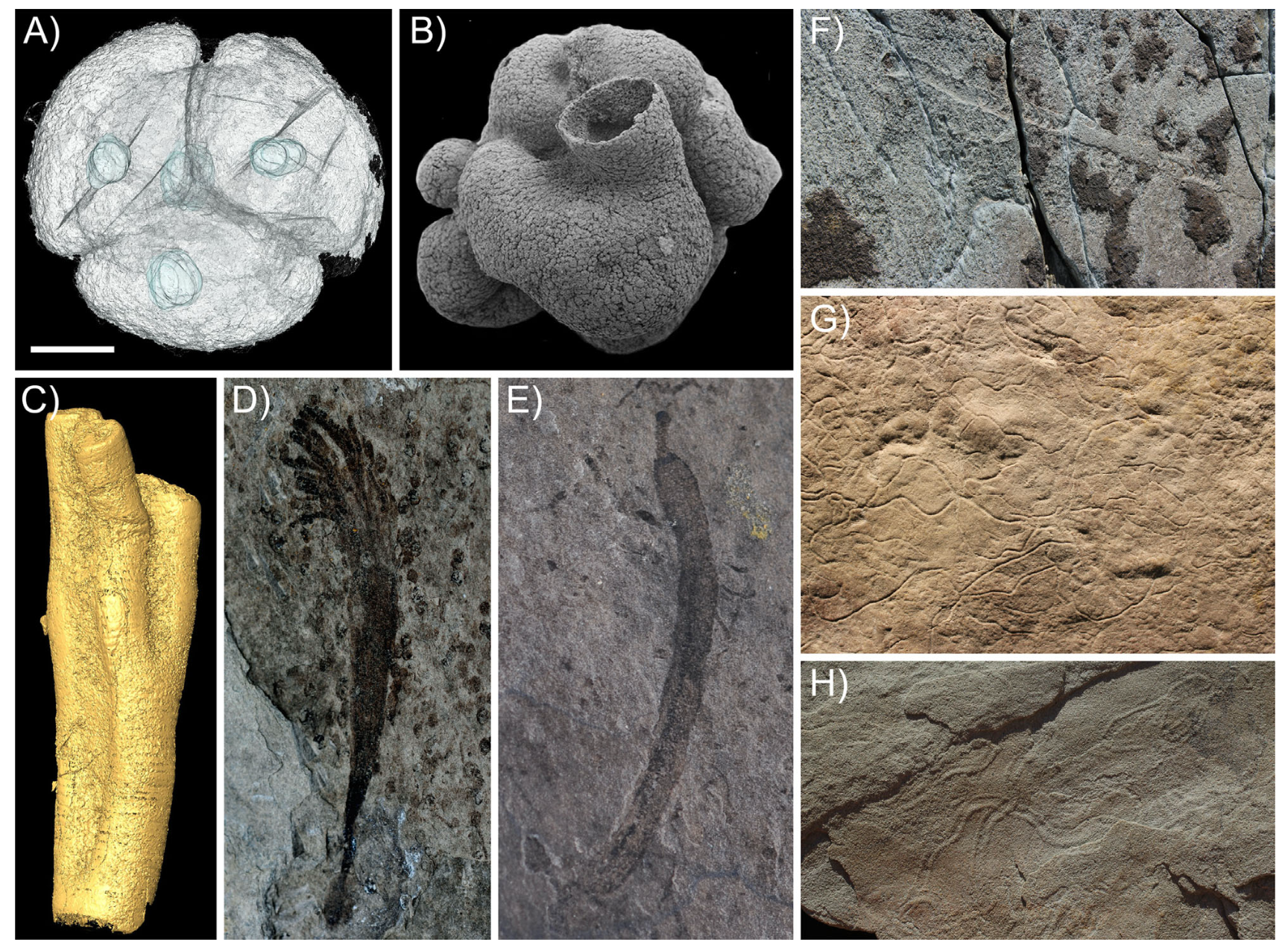
Some putative Ediacaran animal fossils from China and South Australia (from Cunningham et al. 2016). Here is the key from the original caption: “A: Embryo-like fossil Tianzhushania. B: Eocyathispongia. C: Ramitubus. D: Lantiella. E: Xiuningella. F: Putative eumetazoan trace fossil from Mistaken Point, Newfoundland. G: Helminthoidichnites, a putative bilaterian trace fossil. H: Archaeonassa, a putative bilaterian trace fossil.“
Now consider: there are many reports of animal fossils older than the oldest unambiguous body fossils— putative sponge spicules, biomarkers, and wiggles in the sediment suggestive of biological activity (e.g., Cunningham et al. 2016). Most of these are probably inorganic productions. And yet there is a non-zero probability that one or the other is genuinely biological, and beyond that, genuinely animal. Would-be clock-makers must ultimately decide what to do about this.
The traditional response has been to adopt conservative maximum constraints, indicating a high degree of uncertainty about the real age of clades. The approach makes sense when the goal of the clock-maker is “to integrate over uncertainty” (in particular, uncertainties stemming from paleontological disagreements about the interpretation of fossils, and inferences about clade age derived from these interpretations). Yet Carlisle et al. took a different tack: “Here, we seek to minimize this uncertainty, in particular, by departing from the practice of establishing maxima such that they encompass the variable quality of fossil evidence and the broad diversity of interpretations of clade age that are based on it” (Carlisle et al. 2024, 2). The reason was that their goal was not to integrate over uncertainties about the interpretation of the fossil record (producing, in effect, a maximally conservative timescale). It was rather, “to establish an integrative timescale for animal diversification that is based on a critical evaluation of the fossil record and its interpretations rather than an historical consensus view” (2).
Carlisle et al.’s results
What does it mean to “establish an integrative timescale for animal diversification… based on a critical evaluation of the fossil record”? In a word: to produce the best possible estimate of the actual ages of clades by excluding low probability interpretations from the analysis. An example of a low probability interpretation is the suggestion that certain sterol molecules found in rocks of Cryogenian age [> 635 Mya] indicate the presence of sponges. This is a low probability interpretation since it is now known that these molecules form “through the geological methylation of C29 sterols of chlorophyte algae, the dominant eukaryotes at that time” (Nettersheim et al. 2019). So, most likely, the presence of putative sponge “biomarkers” reflects only the presence of algae, not sponges at all.
Could sponges have produced the sterol signature? Yes— but they probably didn’t. So, on the aggressive approach employed by the Carlisle et al. group, this possibility is discounted. It amounts to accepting a trade-off. On the one hand, accommodating the possible truth of low probability interpretations will inflate divergence estimates; on the other, discounting them may produce artificially young estimates if one or more of these interpretations turn out to be true. Neither approach is objectively better than the other. They are simply different things: a best guess, on the one hand, a maximally conservative estimate on the other.
Recognizing this distinction is part of a “new philosophy of molecular clock dating.” But it is not the whole thing. Another component of concerns the question of what molecular clocks are, and how they relate to the fossil record.
Glimpses of a new philosophy of molecular clock dating
To the extent that there was an older “philosophy of molecular clocks,” it was organized around the idea that molecular clocks provide an independent line of evidence bearing on the timing of evolutionary events. Of course, everyone knew that molecular clocks were calibrated using fossils: Wray and colleagues, as we’ve said, calibrated their clock using vertebrate genes and fossils, which together permitted a rate of molecular change to be calculated for each gene. But the fossils used for calibration did not bear, evidentially, on the events of interest, which for Wray et al. were about the divergence of basal animal groups. Instead, they were selected because they permitted a rate of molecular change to be calibrated, which could then be extrapolated to deeper parts of the tree.
Newer Bayesian methods have changed this situation. In the specification of minimum constraints, in particular, calibrations are supplied by the very fossil record of the clades under scrutiny. Consequently, there is no question of treating molecules and fossils as independent lines of evidence bearing on the same problem. They are entangled in the very guts of the clock.*
[* The Wray et al. study was truly independent of the fossil record of invertebrates, since nothing about that fossil record constrained the estimates of clade age produced by the clock. Not so Erwin et al. (2011) and Carlisle et al. (2024). Especially the Carlisle et al. study, with its aim of establishing “an integrative timescale for animal diversification… based on a critical evaluation of the fossil record,” hinged on the interpretation of fossil biotas directly relevant to assessing the age of major clades like the metazoans.]
What, then, is a molecular clock? A method for dating the origin of clades, to be sure. But current Bayesian clocks are also a means of “rationalizing disparate paleontological, molecular, and phylogenetic data, their interpretations, [and] evolutionary models, to obtain a holistic evolutionary timescale” (Carlisle et al. 2024, 9). These clocks are not independent lines of evidence bearing on the ages of clades. They are a kind of integrative technology that scaffolds interpretations of the fossil record based on several lines of evidence and a set of coordinating assumptions. As such, they do not— and arguably cannot— conflict with fossil evidence. What molecular clocks (can) conflict with are interpretations of the fossil record based on fossil evidence: in particular, interpretations that “read” the fossil record literally, or very nearly so. But they do not conflict with fossil evidence as such, because fossil evidence is an integral part of a molecular clock analysis.
None of these points are very controversial. And yet it is striking that molecular clock methods are not usually conceptualized as providing a means of interpreting the fossil record. Typically, when they are discussed in relation to the fossil record, the implied relationship is adversarial: it is rocks versus clocks, in a battle for all the marbles. And why not? Either animals originated around 550 million years ago (as a literal interpretation of the fossil record might suggest), or they originated >800 million years ago— both of these cannot be true. But this is really a battle over interpretations, which use fossil evidence in different ways, and accord it a different status. It is not “rocks versus clocks”; it is the fossil record alone versus a scaffolded view that integrates fossil and molecular evidence to produce a holistic timescale of evolutionary events. Antagonism is an outdated idiom for discussing the relationship between molecular and fossil evidence.
Three lessons
Three more general lessons can be extracted from the preceding discussion. To begin, the new philosophy of molecular clocks highlights the importance, in the historical sciences, of “integrative technologies”: methods for rationalizing disparate lines of evidence (together with interpretations and models), which permit investigators to acquire a holistic view of a complex event or phenomenon. Previously, philosophers have stressed the importance of obtaining evidence that “relies on causally independent processes of trace generation and on conceptually independent detection techniques and inference-warranting bodies of background knowledge” (Chapman and Wylie 2016, 159). This is surely important. But integration is important too, especially when lines of evidence have potentially complementary strengths and limitations. As a recent paper puts it, “The fossil record supplies a direct record of past diversity that frequently includes character combinations, ecological associations, and distributions that are not inferable from analyses of recent taxa alone” (Hopkins et al. 2018, 562). However, “the molecular record can offer insight on what taxa might be missing from the fossil record, especially for certain time periods and environments” (563). The two records thus complement each other: and this means they can be leveraged simultaneously, provided the appropriate scaffolding is in place. This sort of practice warrants more philosophical attention.
A second lesson concerns negative evidence. Although often poo-pooed in philosophical discussions of evidential reasoning, negative evidence features in many historical inferences, to a perhaps surprising degree (Wallach 2019; Brewer 2025). And yet the situation is not as problematic as it seems.
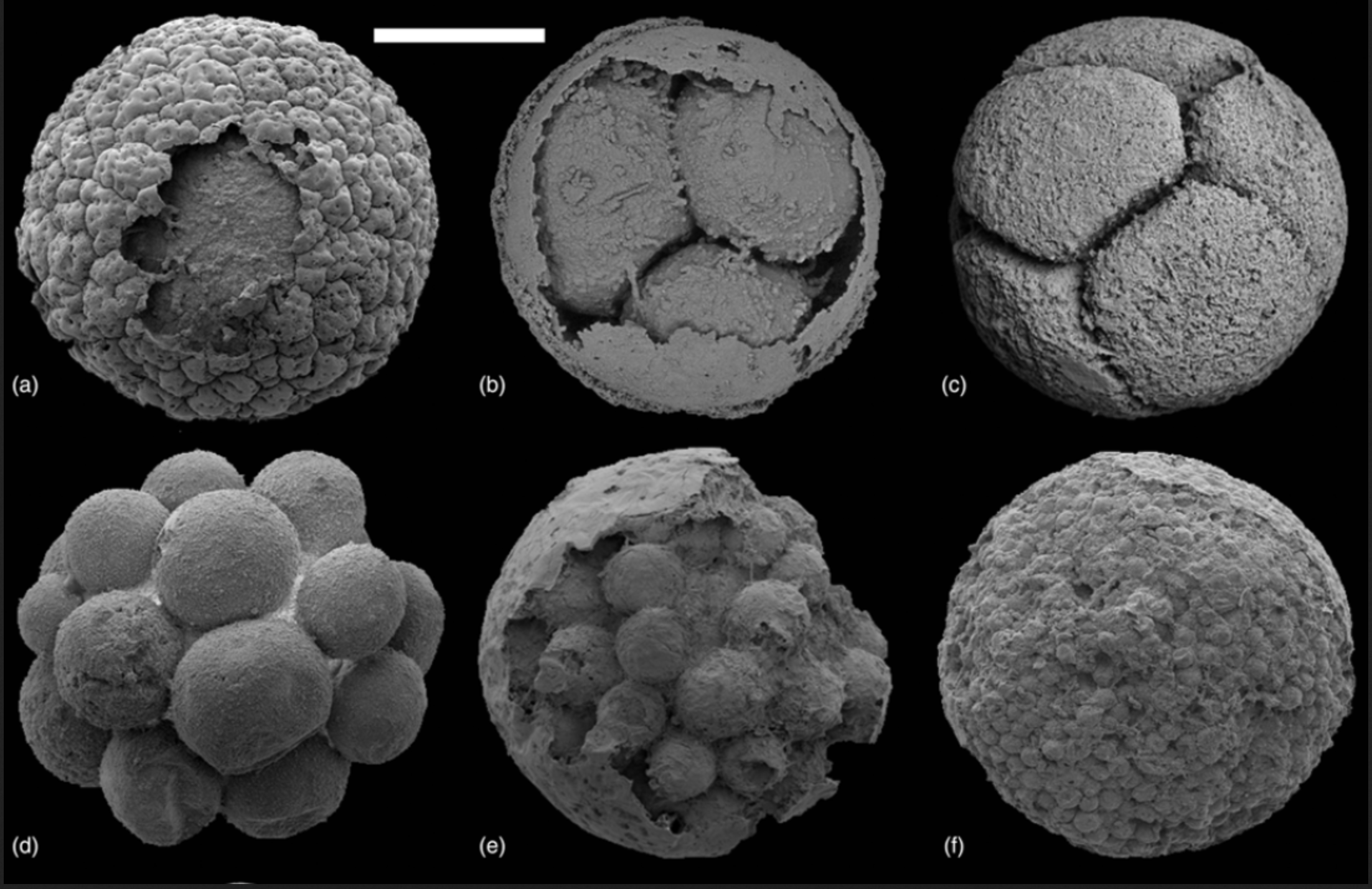
Consider this. In their 2024 study, Carlisle and colleagues base the maximum constraint on the age of crown group Eumetazoa on the Weng’an biota of South China. This biota, which includes three-dimensionally preserved microfossils, has been interpreted as containing animal embryos. Yet as Carlisle et al. put it, “the weight of evidence now favors a non-metazoan holozoan affinity [for the embryo-like structures].” Why then use it as a calibration point? Because the Weng’an location is very likely to have preserved crown group eumetazoans had they been present in the oceans at that time; so the fact that the Weng’an location apparently lacks eumetazoans is strong evidence that eumetazoans had yet to evolve.
Embryo-like fossils from the Weng’an biota (image from Cunningham et al. 2017)
What this shows is that the absence of evidence can be quite powerful evidence of absence under the right circumstances. Consider a second example. A recent study has tried to make sense of the absence of a fossil record for early animals by studying the distribution of fossilization processes over time (Anderson et al. 2023). The authors note that most early animals are preserved as carbonaceous remains in fully marine, fine-grained, siliciclastic rocks (“Burgess Shale-type preservation”). However, in the few assemblages characterized by this mode of preservation dated to 789 Ma or earlier, there is no evidence for metazoans.* They conclude that “taphonomic evidence argues for a maximum constraint of 788.72 ± 0.29 Ma on crown animal [metazoan, not eumetazoan] antiquity” (Anderson et al. 2023, 1067). By characterizing the distribution of fossilization processes over time, and by leveraging background knowledge about the kinds of processes likely to preserve metazoans, scientists can make inference about the likely presence or absence of animal ancestors in fossil biotas.
[* This study is philosophically aligned with the Erwin et al. (2011) study, which sought to provide a conservative estimate for the ages of groups, as opposed to the more aggressive Carlisle et al. (2024) study.]
The third lesson is specific to paleontology. In his 2012 book, Rereading the Fossil Record, David Sepkoski identified three strategies that paleobiologists developed during the 1970s and ‘80s to cope with the shortcomings of the fossil record. The first he termed “literal rereading,” and involved interpreting the fossil record at (more or less) face value. The second he termed “idealized rereading,” and involved simulating the history of life in silico (basically ignoring the fossil record except for the purpose of comparing its broad patterns to those generated by computer simulations). The third he termed “generalized rereading,” and involved assembling large datasets for the purpose of framing “statistical generalizations… about patterns in life’s history” (Sepkoski 2012, 4). Then, after the 1980s, more strategies came online. For example, one of us has previously discussed how paleobiologists began using models of sedimentary accumulation to diagnose and correct for stratigraphic overprint, especially at local and regional scales (Dresow 2023). This, he suggested, was a distinct approach to “rereading the fossil record,” with particular application to field-based research.
We can now add another approach: the use of molecular clocks to produce integrative timescales for evolutionary events. The approach is limited to Bayesian clock methods, which integrate genomic data with fossil-based time constraints, while also accommodating several independent sources of uncertainty. But it is broad enough to encompass both conservative and aggressive approaches to fossil calibration, which produce different outputs: maximally conservative estimates of clade age versus “best guess” integrative timescales, respectively.
Again, there is something unfamiliar about regarding molecular clocks as a means of interpreting the fossil record. The trope of “rocks versus clocks” is not easily forgotten, and is reinforced by an apparently irresistible tendency to discuss the relationship in adversarial terms. (Consider the title of a 2016 paper: “can molecular clocks and the fossil record be reconciled?”) Yet this is so much historical baggage. Rocks and clocks do not conflict. What conflict are interpretations of the fossil record that either do or do not seek to integrate over multiple sources of information and uncertainty. To use Sepkoski’s language, “literal rereading” is at odds with an approach to the fossil record that uses Bayesian methods to obtain integrative evolutionary timescales. But literal rereading is now generally regarded with suspicion; so this tension comes to rather little.
For an enjoyable public discussion of molecular clocks applied to early animal evolution, see the embedded video (especially Phil Donoghue’s presentation, beginning at 12:10, and the Q&A, especially 1:23:25)
References
Anderson, R. P., Woltz, C. R., Tosca, N. J., Porter, S. M., and D. E. G. Briggs. 2023. Fossilisation processes and our reading of antiquity. Trends in Ecology and Evolution 38:1060–1071.
Ayala, F. J., Rzhetsky, A. and F. J. Ayala. 1998. Origin of the metazoan phyla: molecular clocks confirm paleontological estimates. Proceedings of the National Academy of Science, United States of America 95:606–611.
Benton, M. J. 2009. The fossil record: biological or geological signal? in D. Sepkoski and M. Ruse, eds., The paleobiological revolution: essays on the growth of modern paleontology. Chicago: University of Chicago Press, 43–59.
Brewer, M. forthcoming. “When should absence of evidence be evidence of absence? A case study from paleogeology.” Philosophy of Science. doi:10.1017/psa.2025.10125.
Bromham, L., A. Rambaut, R. Fortey, A. Cooper, and D. Penny. 1998. Testing the Cambrian explosion hypothesis using a new molecular dating technique. Proc Natl Acad Sci U S A, 95:12386–12389.
Carlisle, E., Yin, Z., Pisani, D., and P. C. J. Donoghue, “Ediacaran origin and EdiacaranCambrian diversification of Metazoa,” Science Advances 10 (2024), eadp7161.
Chapman, R. and Wylie, A. 2016. Evidential reasoning in archaeology. London: Bloomsbury.
Cunningham, J. A., Liu, A. G., Bengtson, S. and P. C. J. Donoghue. 2016 The origin of animals: can molecular clocks and the fossil record be reconciled? BioEssays 39: 1600120.
Dohrmann, M. and G. Wörheide 2017. Dating early animal evolution using phylogenomic data. Scientific Reports 7:3599.
Dos Reis, M., Donoghue, P. C. J., and Z. Yang. 2016. Bayesian molecular clock dating of species divergences in the genomics era. Nature Reviews Genetics 17:71–80.
Dos Reis, M. Thawornwattana, Y., Angelis, K. Telford, M. J., Donoghue, P. C. J., and Y. Ziheng Yang. 2015. Uncertainty in the timing of the origin of animals and the limits of precision in molecular timescales. Current Biology 25:2939–2950.
Dresow, M. 2023. Biased, spasmodic and ridiculously incomplete: sequence stratigraphy and the emergence of a new approach to stratigraphic complexity in paleobiology, 1973–1995. Journal of the History of Biology 56:419–454.
Erwin, D. H. 1989. Molecular clocks, molecular phylogenies and the origin of phyla. Lethaia 22 (1989), 251–257.
Erwin, D. H., Laflamme, M. Tweedt, S., Sperling, S. A., Pisani, D. and K. Peterson. 2011. Cambrian conundrum: early divergence and later ecological success in the early history of animals. Science 334:1091–1097.
Hopkins, M., Bapst, D. W., Simpson, C. and R. C. M. Warnock. 2018. The inseparability of sampling and time and its in attempts to unify the molecular and fossil records. Paleobiology 44:561–574.
Knoll, A. H. 2003. Life on a Young Planet: The First Three Billion Years of Evolution on Earth. Princeton: Princeton University Press.
Nash, J. M. 1995.“When life exploded. TIME, December 4, 1995.
Nettersheim, B. J. et al. 2019. Putative sponge biomarkers in unicellular Rhizaria question an early rise of animals. Nature Ecology & Evolution 3:577–581.
Runnegar, B. 1982. A molecular-clock date for the origin of the animal phyla. Lethaia 15:199–205.
Sepkoski, D. 2012. Rereading the fossil record: the growth of paleobiology as an evolutionary discipline. Chicago: University of Chicago Press.
Tiley, G. P., Poelstra, J. W., dos Reis, M., Yang, Z. and A. D. Yoder. Molecular clocks without rocks: new solutions for old problems. Trends in Genetics 36:845–856.
Valentine, J. W., Erwin, D. H., and D. Jablonski. 1996. Developmental evolution of metazoan body plans: the fossil evidence,” Developmental Biology 173:373–381.
Wallach, E. 2019. Inference from absence: the case of archaeology. Palgrave Communications. https://doi.org/10.1057/s41599-019-0307-9.
Wang, D. Y.-C., S. Kumar, and S. B. Hedges. 1999. Divergence time estimates for the early history of animal phyla and the origin of plants, animals, and fungi. Proceedings of the Royal Society, Part B 266:163–171.
Wray, G. A., Levinton, J. S., and L. H. Shapiro. 1996. Molecular evidence for deep Precambrian divergences among metazoan phyla,” Science 274:568–573.